Overview
We design new and optimize existing enzymes using computational and experimental tools. Ultimately, we are interested in understanding the fundamental biophysical bases of enzyme function and use this knowledge to engineer enzymes for applications including biodegradation of pollutants and toxins, stabilization of enzymes for harsh environments, extension of enzymatic chemistries beyond what biology has explored, and the design of enzymatic catalytic therapeutics. Our work spans collaborations with academic and industrial partners, including the SaBRe (Sargassum BioRefinery) consortium, which focuses on enzyme engineering for marine biomanufacturing.
Designing Enzyme Selectivity
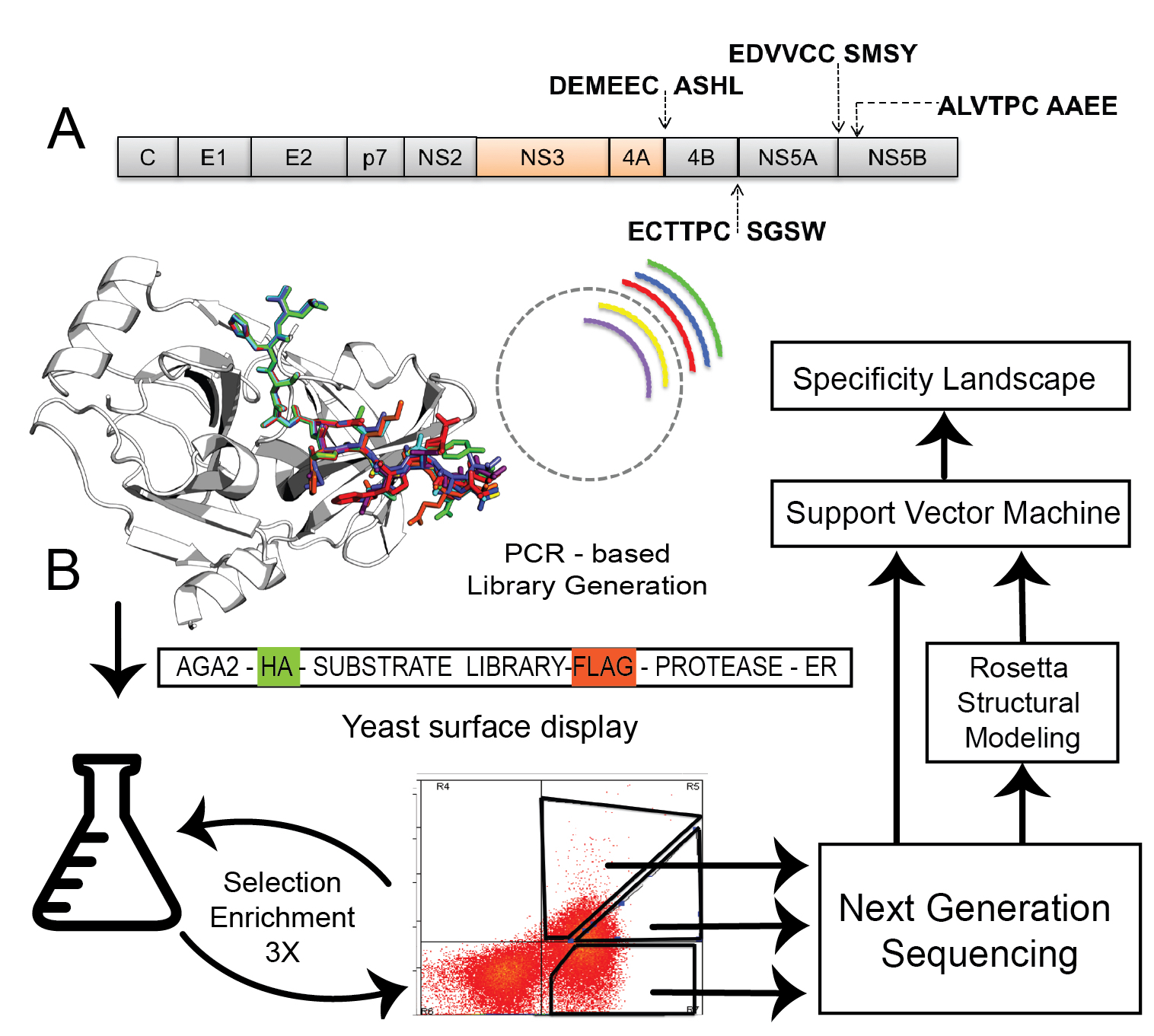
Figure adapted from Lu, Lubin, Sarma et al., Proc. Natl. Acad. Sci. USA 120:e2303590120 (2023).
Enzymes achieve their remarkable rate accelerations in part by being highly selective for a particular substrate or stereochemical outcome. We are interested in reshaping that selectivity, expanding it to new substrates, or directing it toward unnatural stereochemistries, with applications in catalytic therapeutics, biopolymer synthesis, and asymmetric chemistry.
Protease specificity for catalytic drugs
Site-selective proteolytic cleavage is a ubiquitous post-translational modification, and proteases with "dialed-in" substrate selectivities would be ideal catalytic drugs, capable of irreversibly neutralizing target substrates such as viral coat proteins. We have developed a mechanism-guided biophysical framework, more recently extended with a structure-aware graph convolutional network, that allows design for both positive and negative substrate specificity, coupled with high-throughput experimental testing.
Looking ahead, we envision designed proteases as a selective catalytic module in protein-editing systems, paired with custom-designed peptide-binding domains as the recognition module. Combining these two designable components delivers a one-two punch for selective protein targeting.
Key papers: Pethe 2019, Lu 2023, Pethe 2017.
Protease-catalyzed polymer and peptide synthesis
In collaboration with the Gross lab (Rensselaer/Rutgers), we use computationally guided proteases as catalysts for the synthesis of biodegradable peptide polymers and oligo-amino-acid chains. Computational modeling guides substrate redesign and provides specificity insights that drive iterative improvements in polymer chemistry.
Key papers: Black 2025, Yang 2020.
Designing stereoselective and stereodivergent biocatalysts
Building on a long-standing collaboration with the Fasan lab, we have developed computationally designed enzyme variants that catalyze stereodivergent transformations. Our 2026 paper reports the first generalist cyclopropanases capable of producing the full set of cyclopropane stereoisomers from a broad range of olefin substrates with high diastereo- and enantioselectivity.
Key papers: Shen 2026, Ren 2022, Nam 2022, Aggarwal 2022.
Protein and Enzyme Stabilization
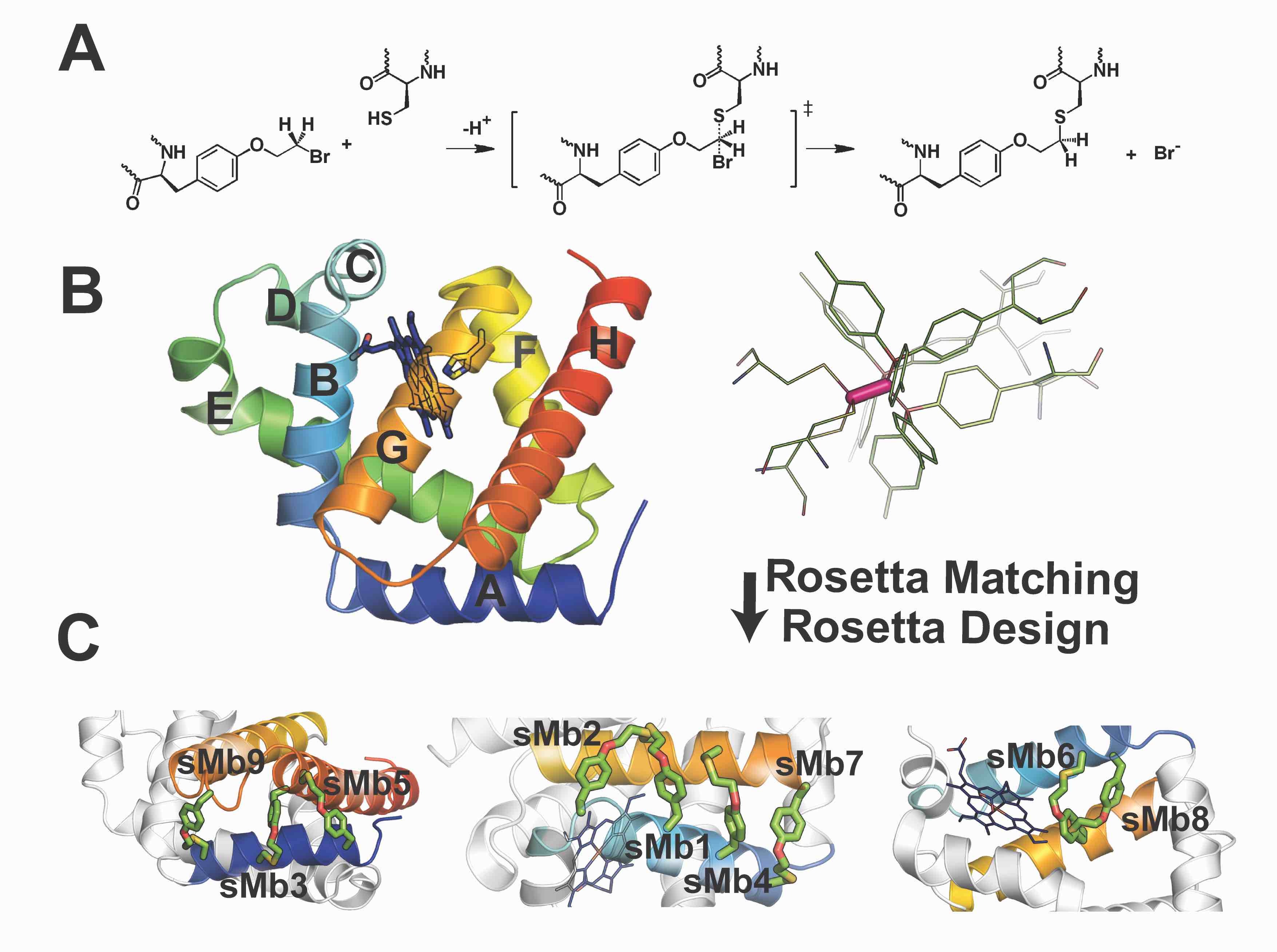
Figure adapted from Moore, Zorine, Hansen, Khare, & Fasan, Proc. Natl. Acad. Sci. USA 114:12472–12477 (2017).
Currently available methods for protein stabilization, such as directed evolution and consensus mutagenesis, can be labor-intensive and often involve extensive amino-acid substitutions that may impair activity or selectivity. We are developing computational methods that combine physics-based modeling and AI/ML approaches to identify minimally invasive modifications that stabilize proteins and enzymes against thermal denaturation and nonaqueous environments.
Our physics-based approach, Rosetta-guided protein stapling (R-GPS), uses structure-based modeling to introduce covalent "staples" into a protein scaffold via genetically encodable noncanonical amino acids. Stapled variants of a stereoselective cyclopropanation biocatalyst show greatly increased thermostability and robustness to high concentrations of organic cosolvents.
We complement these physics-based methods with AI/ML approaches that learn structural and sequence features predictive of protein stability, robustness to mutation, and tolerance to nonaqueous environments. These approaches are increasingly integrated with our physics-based design pipeline.
We are also part of the SaBRe (Sargassum BioRefinery) consortium, a Schmidt Sciences-funded virtual research center that aims to transform Sargassum seaweed into a renewable feedstock for biomanufacturing. In this context, we are engineering enzymes that operate stably in marine conditions and tolerate the salinity, variable temperatures, and complex substrate mixtures characteristic of seaweed-derived feedstocks.
Key papers: Moore 2017, Iannuzzelli 2022.
Stimulus-Responsive Enzymes and Enzyme Assemblies
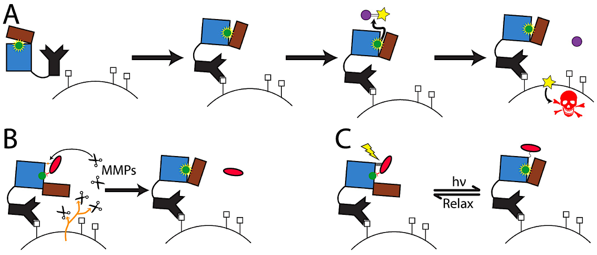
Figure adapted from Blacklock, Yachnin, Woolley, & Khare, J. Am. Chem. Soc. 140:14–17 (2018).
Many therapeutic and biosensing applications require enzymes whose activity can be precisely controlled in space and time, or assembled at high local concentration to coordinate cascades of reactions. We design enzymes that respond to defined stimuli (such as disease-specific proteases or visible light), and supramolecular assemblies that organize component enzymes for enhanced catalysis or sensing.
Smart enzymes gated by chemical or optical stimuli
Conventional chemotherapy is limited by the lack of spatial and temporal control over the action of the toxic drug. Computationally designed enzymes that are constitutively inactive but activatable by a tumor-specific stimulus (such as MMP-2 protease) or by light (via attached azobenzene dyes or genetically encoded photoswitchable amino acids) are expected to overcome these limitations. We have obtained ~10X switches straight from computational design and developed high-throughput screening approaches to improve them, including designed proenzymes activated by tumor proteases. We collaborate with the Deiters Lab (Univ. of Pittsburgh) on optical control of protein function.
Key papers: Yachnin & Khare 2017, Blacklock 2018, Yasuike 2019, Dolan & Khare 2022, Yachnin 2022, Janosko 2023.
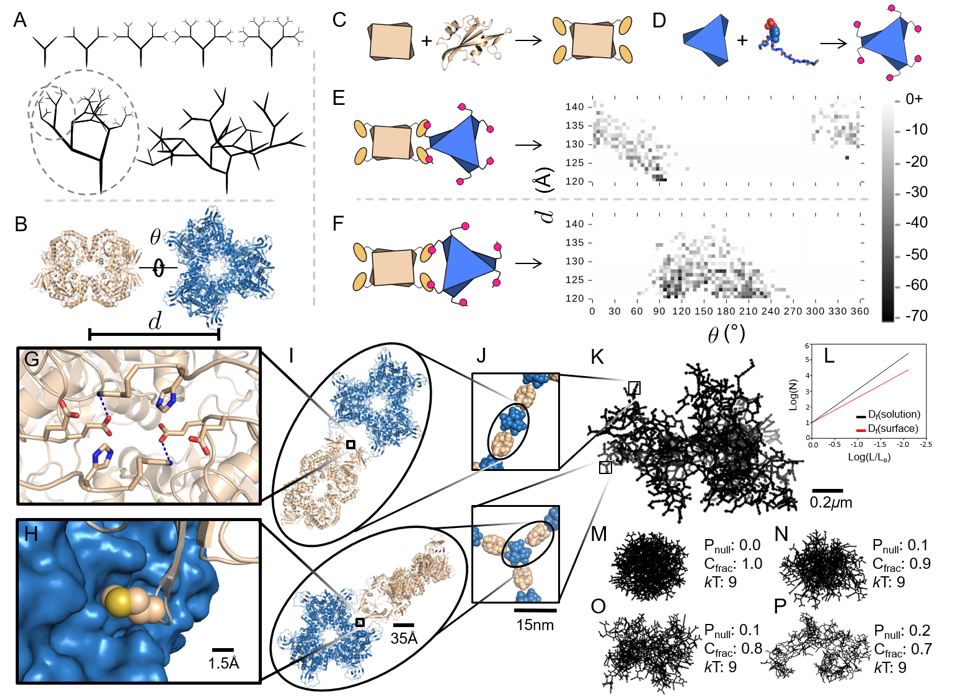
Figure adapted from Hernández et al., Nature Chemistry 11:605–614 (2019).
Supramolecular enzyme assemblies
Enzymatic processes in nature are spatially organized. To similarly organize synthetic enzymatic pathways and develop efficient biosensors (which benefit from high surface-area-to-volume ratios), we have developed a modular design approach that allows construction of supramolecular assemblies in response to chemical or optical stimuli. We have built fractal supramolecular topologies that organize component enzymes into hyperbranched dendritic architectures. Although fractals are ubiquitous in nature, our studies represent the first instance of designing such topologies via protein self-assembly. We are now applying this technology to biodegradation, taking advantage of the high surface-area:volume ratio for sponge-like substrate uptake.
Key papers: Hernández 2019, Hansen & Khare 2020, Yang 2017.
Novel Computational Design Methods
New computational design methods underpin all of our research, spanning physics-based modeling, AI/ML-driven design, and the integration of the two.
Physics-based modeling and design with unnatural amino acids
In Rosetta, we have introduced a Potts-model-based modeling and design algorithm, methods for designing nested (domain-inserted) proteins, and methods for designing metalloproteins and other proteins with genetically encodable noncanonical (unnatural) amino acids. We pursue force-field improvements jointly with the Case lab (Rutgers), reconciling Rosetta and Amber energy evaluations.
Custom neural networks and language models for protein design
We develop custom neural networks and language models tailored to protein-design problems, including graph attention networks and graph convolutional networks for predicting functional sequences and substrate specificity at protein interfaces, and constrained generative diffusion approaches that enforce hard structural constraints during protein design.
Covalent protein design
We design covalent protein architectures, including internal staples and crosslinks that stabilize protein scaffolds, and covalent peptide modifiers that selectively engage target proteins for therapeutic applications.
Key papers: Khare 2012, Tinberg & Khare 2013, Mills 2013 (UAA metalloprotein), Blacklock 2018 (nested proteins), Leman 2020 (Rosetta methods), Rubenstein 2018 (Amber/Rosetta), Lu 2023 (graph CNN), Ash 2025 (graph attention), Christopher 2026 (constrained diffusion), Moore 2017 (stapling), Emanuelson 2025 (covalent modifiers).